Journal: bioRxiv
Article Title: Functional profiling of KCNE1 variants informs population carrier frequency of Jervell and Lange-Nielsen syndrome type 2
doi: 10.1101/2025.03.28.646046
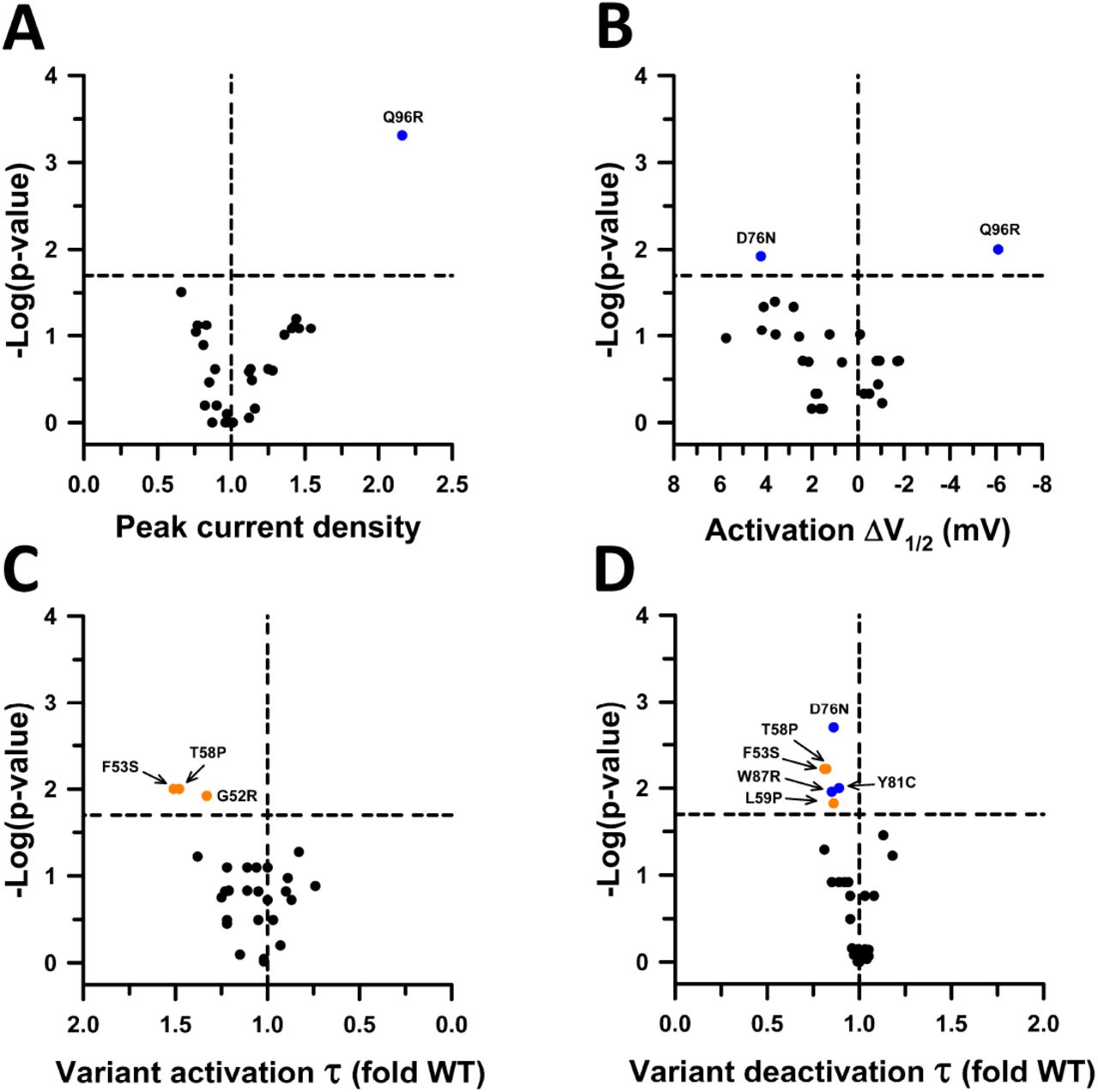
Figure Lengend Snippet: Functional properties of KCNE1 variants in the heterozygous state. ( A ) Volcano plots of mean JNJ-303 sensitive peak whole-cell current density measured at +60 mV from CHO-E1 cells co-expressing WT KCNQ1 with each KCNE1 variant. Data are displayed as fold divergence from WT channels recorded in parallel. Only variants with peak current density significantly different from WT ( P < 0.02, horizontal dashed line) are labeled. Values to the right or left of the vertical dashed line (normalized WT value) represent current density larger or smaller than WT, respectively. ( B ) Volcano plots of mean difference (ΔV½ in mV) between the averaged V½ for variant and WT channels recorded in parallel. Only variants with peak current density significantly different from WT (P<0.02, horizontal dashed line) are labeled. Values to the right or left of the vertical dashed line indicate hyperpolarized or depolarized activation V½, respectively. ( C ) Volcano plots of mean activation time constants (τ) determined for each heterozygous KCNE1 variant plotted as the ratio to the averaged activation time constant for WT channels recorded in parallel. Values to the right or left of the vertical dashed line (no difference from WT) indicate faster or slower activation, respectively. ( D ) Volcano plots of mean deactivation time constants (τ) determined for each heterozygous KCNE1 variant plotted as the ratio to the averaged deactivation time constant for WT channels recorded in parallel. Values to the right or left of the vertical dashed line indicate slower or faster deactivation, respectively. Symbols colors are defined in previous figures. The number of recorded cells for each variant and individual P -values are presented in Supplemental Dataset 2 . Scatter plots of individual data are presented in Supplemental Figure S8 .
Article Snippet: Full length cDNA encoding WT KCNQ1 (GenBank accession number AF000571 ) was engineered in the mammalian expression vector pIRES-CyOFP1 (pIRES-CyOFP1-KCNQ1, AddGene #173162).
Techniques: Functional Assay, Expressing, Variant Assay, Labeling, Activation Assay